
Allowing gene transfers doesn't make life easier for inferring orthology and paralogy
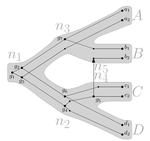
Consistency of orthology and paralogy constraints in the presence of gene transfers
Abstract
Recommendation: posted 16 February 2022, validated 21 February 2022
Holland, B. (2022) Allowing gene transfers doesn't make life easier for inferring orthology and paralogy. Peer Community in Mathematical and Computational Biology, 100009. https://doi.org/10.24072/pci.mcb.100009
Recommendation
Determining if genes are orthologous (i.e. homologous genes whose most common ancestor represents a speciation) or paralogous (homologous genes whose most common ancestor represents a duplication) is a foundational problem in bioinformatics. For instance, the input to almost all phylogenetic methods is a sequence alignment of genes assumed to be orthologous. Understanding if genes are paralogs or orthologs can also be important for assigning function, for example genes that have diverged following duplication may be more likely to have neofunctionalised or subfunctionalised compared to genes that have diverged following speciation, which may be more likely to have continued in a similar role.
This paper by Jones et al (2022) contributes to a wide range of literature addressing the inference of orthology/paralogy relations but takes a different approach to explaining inconsistency between an assumed species phylogeny and a relation graph (a graph where nodes represent genes and edges represent that the two genes are orthologs). Rather than assuming that inconsistencies are the result of incorrect assessment of orthology (i.e. incorrect edges in the relation graph) they ask if the relation graph could be consistent with a species tree combined with some amount of lateral (horizontal) gene transfer.
The two main questions addressed in this paper are (1) if a network N and a relation graph R are consistent, and (2) if – given a species tree S and a relation graph R – transfer arcs can be added to S in such a way that it becomes consistent with R?
The first question hinges on the concept of a reconciliation between a gene tree and a network (section 2.1) and amounts to asking if a gene tree can be found that can both be reconciled with the network and consistent with the relation graph. The authors show that the problem is NP hard. Furthermore, the related problem of attempting to find a solution using k or fewer transfers is NP-hard, and also W[1] hard implying that it is in a class of problems for which fixed parameter tractable solutions have not been found. The proof of NP hardness is by reduction to the k-multi-coloured clique problem via an intermediate problem dubbed “antichain on trees” (Section 3). The “antichain on trees” construction may be of interest to others working on algorithmic complexity with phylogenetic networks.
In the second question the possible locations of transfers are not specified (or to put it differently any time consistent transfer arc is considered possible) and it is shown that it generally will be possible to add transfer edges to S in such a way that it can be consistent with R. However, the natural extension to this question of asking if it can be done with k or fewer added arcs is also NP hard.
Many of the proofs in the paper are quite technical, but the authors have relegated a lot of this detail to the appendix thus ensuring that the main ideas and results are clear to follow in the main text. I am grateful to both reviewers for their detailed reviews and through checking of the proofs.
References
Jones M, Lafond M, Scornavacca C (2022) Consistency of orthology and paralogy constraints in the presence of gene transfers. arXiv:1705.01240 [cs], ver. 6 peer-reviewed and recommended by Peer Community in Mathematical and Computational Biology. https://arxiv.org/abs/1705.01240
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
no declaration
Evaluation round #1
DOI or URL of the preprint: https://arxiv.org/abs/1705.01240
Version of the preprint: 4
Author's Reply, 02 Feb 2022
Decision by Barbara Holland, posted 10 Jan 2022
I am pleased to have (finally) managed to get two expert reviews for this paper. Apologies for how long this took!
You'll see that they both have constructive suggestions that you may like to consider in a revised version.
I haven't delved tinto the technical detail but noticed a few small things.
Page 1
Orthology and paralogy relations are often inferred by methods based on gene sequence similarity, which yield a graph depicting the relationships between gene pairs.
->
Orthology and paralogy relations are often inferred by methods based on gene sequence similarity that yield a graph depicting the relationships between gene pairs.
I always get confused about when to use which and that but here I think that is better (i.e. an essential clause)
Vertical descent with modification (speciation) constitutes only part of the events shaping a gene history;
I'd say that vertical descent with modification is different from speciation, i.e. it's evolution along an edge whereas speciation is a splitting event.
page 3
The authors ask, given a reconciled gene tree G that displays a given of relations, whether there is a species network N that be reconciled with G.
->
The authors ask, given a reconciled gene tree G that displays a given of relations, whether there is a species network N that can be reconciled with G.
page 7 (last sentence)
missing a reference
It is worth mentioning the question studied in ??
Reviewed by anonymous reviewer 1, 09 Sep 2021
Reviewed by anonymous reviewer 2, 03 Jan 2022
The paper presents a new approach to several problems on the homology relations in the gene tree-network reconciliation approach. From the mathematical and algorithmic point of view, the results are correct and sound. In general, the submitted article presents interesting algorithmic and computational complexity results. However, it is formally quite technical and requires some effort to follow. My general recommendation is positive, but I think the article requires revision. Below, I present more detailed remarks on the submitted contribution.
Perhaps, the most tricky elements of the paper are definitions with plenty of symbols and sometimes confusing usage of notions (see comment on Sect. 5). That's both on the level of the definitions and examples. Therefore, I recommend providing some better illustrations with explanations.
Recommendations:
1. Instead of writing a paragraph with exemplary alpha mapping (in pg. 2, which seems to contain mistakes), I recommend providing a picture of G embedded into N with explanations. It would be beneficial in understanding the concept of gene tree-network reconciliations. The current approach might be too difficult for a reader without experience in such approaches.
2. Also, the labeling e^* should be explained directly in Definition 2.
3. In Fig. 2, a comment should be on the presence of edge (c2,b2), since the edge seems not to represent an orthology relation from the exemplary reconciliation of G and N (which is confusing given the definition of orthology relation; however, it is formally correct, since the authors do not claim that R represents the relations from N).
4. Related to the above comment. Pg. 6, Sect. 2.2. Clarify that R is not the orthology graph for N (from Figure) or correct Fig. 1.
5. To be checked on page 6 (in the example):
- in e(alpha_1(b_1)) repeated,
- e(alpha_1(b_2)) missing,
- e(alpha_1(g_5))=S (not T)
- e(alpha_2(g_5))=T missing
6. In Section 5.
Definition 5 is conflicted with Definition 3. If S is a species tree, it is also a network with k=0 transfers. Also, "using k transfers", allows using 0 transfers. Thus, the notion of S-consistency is conflicted with Definition 3, when N is a species tree. The tricky part is that both notions are connected (also in the proof of Lemma 5). A careful reader can understand which definition must be applied, but it took me a while to untangle this issue.
Suggestion: try to avoid using S-consistency and N-consistency, where N and S are defined as a network and a species tree, resp.; Maybe use "species tree-consistency"?
Other comments:
- pg. 11. MWACT instead of ACT (2nd problem definition)
- The conditions on the weight functions are repeated several times (Lemma 3, Lemma 4, proofs, and other parts); I suggest introducing a new notion for the properties and removing the repeated lists.
- pg 6. the last line missing reference ??
- pg 23. 7 line from the top, remove "edge"
- pg 28. 2nd line from the top, LAST subscript;
- pg 28. 3rd line "alpha ... are incomparable" - explain what does it mean in a network
- inconsistent notation of edges: xy or (x,y) in several places
Algorithms.
The presented algorithms are clear and easy to understand Algorithm 2 can be improved by adopting more refined techniques from algorithmic papers on HGT reconciliation (see suggested papers at the end of the review), where the factor O(|V|^2) can be replaced by O(1). Such an update requires the introduction of an additional formula, which for (g,s) returns the minimum cost under the assumption the g is mapped to s', where there is a path from s to s' in N, plus the cost of transfers on the best path from s to s' (note that t(s,s') will be not needed). I leave the decision to the authors on how to incorporate this observation into the results. Such an improvement is not crucial in the contribution (even if the improvement in the polynomial part of FPT algorithm is significant), so a comment would be sufficient.
Please provide space complexity analysis.
Proofs in the appendix.
I also analyzed the proofs in the appendix, focusing on the more demanding and non-trivial proofs on reductions. This part is nicely written and presented but requires the most effort. I did not analyze the proofs of correctness of Algorithms 1 and 2 (the algorithms were easy to follow) and the proof of Theorem 4 (due to reviewing deadlines).
Related work suggestions.
1. The suggested improvement in Alg. 2 is presented in several algorithmic papers on variants of reconciling a gene tree with a species tree with horizontal gene transfer e.g. by Bansal or Mykowiecka (DOI: 10.1109/TCBB.2017.2707083).
2. Modelling reconciliation with transfers: papers on H-trees by Gorecki et. al., which seems the most related to the reconciliation of (G,alpha) with the network N.
3. The question of consistency and existence of reconciliations relates to "reconciliation feasibility problems" which seem to be a simpler version of consistency problems: given sigma mapping and a species tree S, the question is whether there is a gene tree for sigma that reconciles with the S. Also, there are more related questions e.g., on minimizing costs etc. See algorithmic papers by Eulenstein and others.
4. Another feasibility-related paper: see M. Helmuth, 2017.
https://doi.org/10.24072/pci.mcb.100108.rev12









