
Comparing reconciled gene trees in linear time
 based on reviews by Barbara Holland, Gabriel Cardona, Jean-Baka Domelevo Entfellner and 1 anonymous reviewer
based on reviews by Barbara Holland, Gabriel Cardona, Jean-Baka Domelevo Entfellner and 1 anonymous reviewer
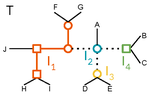
A linear time solution to the Labeled Robinson-Foulds Distance problem
Abstract
Recommendation: posted 24 December 2020, validated 24 December 2020
Scornavacca, C. (2020) Comparing reconciled gene trees in linear time. Peer Community in Mathematical and Computational Biology, 100002. https://doi.org/10.24072/pci.mcb.100002
Recommendation
Unlike a species tree, a gene tree results not only from speciation events, but also from events acting at the gene level, such as duplications and losses of gene copies, and gene transfer events [1]. The reconciliation of phylogenetic trees consists in embedding a given gene tree into a known species tree and, doing so, determining the location of these gene-level events on the gene tree [2]. Reconciled gene trees can be seen as phylogenetic trees where internal node labels are used to discriminate between different gene-level events. Comparing them is of foremost importance in order to assess the performance of various reconciliation methods (e.g. [3]).
A paper describing an extension of the widely used Robinson-Foulds (RF) distance [4] to trees with labeled internal nodes was presented earlier this year [5]. This distance, called ELRF, is based on edge edits and coincides with the RF distance when all internal labels are identical; unfortunately, the ELRF distance is very costly to compute. In the present paper [6], the authors introduce a distance called LRF, which is inspired by the TED (Tree Edit Distance [7]) and is based on node edits. As the ELRF, the new distance coincides with the RF distance for identically-labeled internal nodes, but has the additional desirable features of being computable in linear time. Also, in the ELRF distance, an edge can be deleted if only it connects nodes with the same label. The new formulation does not have this restriction, and this is, in my opinion, an improvement since the restriction makes little sense in the comparison of reconciled gene trees.
The authors show the pertinence of this new distance by studying the impact of taxon sampling on reconciled gene trees when internal labels are computed via a method based on species overlap. The linear algorithm to compute the LRF distance presented in the paper has been implemented and the software —written in Python— is freely available for the community to use it. I bet that the LRF distance will be widely used in the coming years!
References
[1] Maddison, W. P. (1997). Gene trees in species trees. Systematic biology, 46(3), 523-536. doi: https://doi.org/10.1093/sysbio/46.3.523
[2] Boussau, B., and Scornavacca, C. (2020). Reconciling gene trees with species trees. Phylogenetics in the Genomic Era, p. 3.2:1–3.2:23.
[3] Doyon, J. P., Chauve, C., and Hamel, S. (2009). Space of gene/species trees reconciliations and parsimonious models. Journal of Computational Biology, 16(10), 1399-1418. doi: https://doi.org/10.1089/cmb.2009.0095
[4] Robinson, D. F., and Foulds, L. R. (1981). Comparison of phylogenetic trees. Mathematical biosciences, 53(1-2), 131-147. doi: https://doi.org/10.1016/0025-5564(81)90043-2
[5] Briand, B., Dessimoz, C., El-Mabrouk, N., Lafond, M. and Lobinska, G. (2020). A generalized Robinson-Foulds distance for labeled trees. BMC Genomics 21, 779. doi: https://doi.org/10.1186/s12864-020-07011-0
[6] Briand, S., Dessimoz, C., El-Mabrouk, N. and Nevers, Y. (2020) A linear time solution to the labeled Robinson-Foulds distance problem. bioRxiv, 2020.09.14.293522, ver. 4 peer-reviewed and recommended by PCI Mathematical and Computational Biology. doi: https://doi.org/10.1101/2020.09.14.293522
[7] Zhang, K., and Shasha, D. (1989). Simple fast algorithms for the editing distance between trees and related problems. SIAM journal on computing, 18(6), 1245-1262. doi: https://doi.org/10.1137/0218082
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
Evaluation round #2
DOI or URL of the preprint: 10.1101/2020.09.14.293522
Decision by Céline Scornavacca , posted 07 Dec 2020
, posted 07 Dec 2020
Dear authors,
The reviewers are very satisfied with the second version. Thank you for your work, it's a nice paper that I will recommend very gladly!
Before I can post my recommendation, there are three comments from one of the reviewers asking for small changes, and a few comments of my own.
Reviewer's comments :
The definition of island should be improved. For instance, using the term "maximum subtree" is not correct, since there are non-comparable subtrees; maximal would be the right word for it. Moreover, saying that it has a maximum number of edges makes that, for instance, I2 in Figure 3 is no longer an island (since it does not have the maximum number of edges, which is 3, as in I1). Also, to make the sentence more readable, "no internal edge of I is a good edge of T" could be replaced by "all its internal edges are bad edges of T". Finally, I'd still suggest the formulation using connected components: Let \hat T be the forest obtained by removing from T the good edges; then, an island is a connected component of \hat T together with the good edges of T adjacent to this component.
In Definition 1, apart from changing "children" by "neighbour", the authors have kept the sentences describing deletions. I still think that the definition is not correct: They do not want that the neighbours of x become the neighbours of y, but want to add them to the existing neighbours of y. Simply deleting the word "the" would be enough.
[...] Also notice that "bipartitions1" is later refered to as "bipartitions_1". This comment about typography also applies to the operations "Del" and "Ins" in Definition 1.
My comments :
- "While remaining a metric, ELRF turned out to be much more challenging to compute. As a result, only a heuristic could be proposed to compute it."
I would rather say that you proposed a heuristic (an approximate or FPT algorithm would have been possible). What I mean here is that NP-hard does not imply that only heuristics are possible.
- " We denote by L(T) \subseteq V (T) the set of leaves of T, i.e. the set of nodes of T of degree one. In particular, given a set L (let us say taxa or genetic elements), a tree T on L is a tree with leafset L(T) = L."
Nodes and labels are mixed up here : you cannot say " L(T) included in V (T)" if they are labels and you cannot say "Given two unrooted trees T and T0 on the leafset L" if they are leaves. Please clarify.
- Edge contraction Cont(T, {x, y}) is equal to a node deletion Del(T, y, x) deleting y, but contrary to LRF, requires that lambda(x) = lambda(y);
Why not say that your solution makes more sense for what you are doing here because it doesn't require lambda(x) = lambda(y)? For me a reconciliation is node-centered and not edge-centered.
- I suggest you move a paragraph (see attached document).
Could you send me an informal response to these points and post a new version on bioarxiv so that I can make my recommendation?
Thanks,
Céline
Additional request from the managing board
To format your final version of your MS
Mandatory modifications
1- Please make sure that:
-Data are available to readers, either in the text or through an open data repository such as Zenodo (free), Dryad or some other institutional repository. Data must be reusable, thus metadata or accompanying text must carefully describe the data.
-Details on quantitative analyses (e.g., data treatment and statistical scripts in R, bioinformatic pipeline scripts, etc.) and details concerning simulations (scripts, codes) are available to readers in the text, as appendices, or through an open data repository, such as Zenodo, Dryad or some other institutional repository. The scripts or codes must be carefully described so that they can be reused.
-Details on experimental procedures are available to readers in the text or as appendices.
-Authors have no financial conflict of interest relating to the article. The article must contain a "Conflict of interest disclosure" paragraph before the reference section containing this sentence: "The authors of this preprint declare that they have no financial conflict of interest with the content of this article." If appropriate, this disclosure may be completed by a sentence indicating that some of the authors are PCI recommenders: "XXX is one of the PCI Math Comp Biol recommenders."
2- Please make the following changes:
-Add the following sentence in the acknowledgements: "Version 4 of this preprint has been peer-reviewed and recommended by Peer Community In Mathematical and Computational Biology (https://doi.org/10.24072/pci.mcb.100002)"
-If you use bioRxiv to post your preprint, add this latter sentence also in the “revision summary” section of the deposit form of bioRxiv.
Note that this DOI is not the DOI of your article, but the DOI of the recommendation text. The DOI of your article remains unchanged.
3- If not yet done, please send us a picture for which you own the rights that could serve as a thumbnail/illustration for your article on the web site of PCI. It can be a figure of the article.
Optional instructions (we strongly advise you to follow them)
1- We suggest you to remove line numbering from the preprint and put the tables and figures within the text rather than at the end of your MS.
2- Then, we strongly advise you to use the PCI templates (word docx template or latex template) to format your preprint in a PCI style. Here is the links of the templates: https://peercommunityin.org/templates/
→ For word template:
Do not hesitate to modify the template as you want (and send it back to us if you made significant improvements).
-the text to be replaced by your own text starts with XXX, eg XXXXTitle of the article.
-XXXXthe "citeas" → Briand, S., Dessimoz, C., El-Mabrouk, N. and Nevers, Y. (2020) A linear time solution to the labeled Robinson-Foulds distance problem. bioRxiv, 2020.09.14.293522, ver. 4 peer-reviewed and recommended by PCI Mathematical and Computational Biology. doi: https://doi.org/10.1101/2020.09.14.293522
-XXXXthe date of deposit in the preprint server → date of the deposit of the latest version
-XXXXthe surnames and names of the reviewers we sent you → Jean-Baka Domelevo Entfellner, Gabriel Cardona, Barbara Holland and one anonymous reviewer
-XXXXthedoiwesentyou → https://doi.org/10.24072/pci.mcb.100002
-XXXXthe surname and name of the recommender → Céline Scornavacca
-In the acknowledgements, add this sentence → "Version 4 of this preprint has been peer-reviewed and recommended by Peer Community In Mathematical and Computational Biology (https://doi.org/10.24072/pci.mcb.100002)"
-Please be careful to choose the badges “Open Code” and “Open Data” only if appropriate (in addition to the “Open Access” and “Open Peer-Review” badges).
→ For Latex and mode org templates:
Do not hesitate to modify the template as you want (and send it back to us if you made significant improvements).
-main.tex and sample.bib should be filled.
-in main.tex, the recommender’s name is "Céline Scornavacca" and the reviewers’ names are Jean-Baka Domelevo Entfellner, Gabriel Cardona, Barbara Holland and one anonymous reviewer
-In sample.bib, indicate the right version of your preprint. It is version 4
-Preambulemcb.tex should be modified (comment lines 115, 119) to select badges. Please be careful to choose the badges “Open Code” and “Open Data” only if appropriate (in addition to the “Open Access” and “Open Peer-Review” badges).
3- we suggest that you deposit a copy of your MS in zenodo.org and ask for its inclusion in the PCI community (“Communities” section in the deposit form). Indicate the current doi of your MS, if it already has one, in the “doi” section.
I hope this is clear. Do not hesitate to ask for any help if needed.
Once you have made these modifications, you should upload the new version of the article on the preprint server. Please tell us when you have done so. Thanks.
Reviewed by Jean-Baka Domelevo Entfellner , 01 Dec 2020
I am full OK with this second version, the authors have taken into account my suggestions and I believe the manuscript is now greatly enhanced, ready for publication. Thank you.
https://doi.org/10.24072/pci.mcb.100002.rev21Reviewed by Gabriel Cardona, 22 Nov 2020
The authors have addressed all the issues I raised on my first review, most of them in a satisfactory way. Some points I'd like to remark:
I still think that the name "Labeled Robinson Foulds" is misleading even though it has not been used elsewhere.
The definition of island should be improved. For instance, using the term "maximum subtree" is not correct, since there are non-comparable subtrees; maximal would be the right word for it. Moreover, saying that it has a maximum number of edges makes that, for instance, I2 in Figure 3 is no longer an island (since it does not have the maximum number of edges, which is 3, as in I1). Also, to make the sentence more readable, "no internal edge of I is a good edge of T" could be replaced by "all its internal edges are bad edges of T". Finally, I'd still suggest the formulation using connected components: Let \hat T be the forest obtained by removing from T the good edges; then, an island is a connected component of \hat T together with the good edges of T adjacent to this component.
- In Definition 1, apart from changing "children" by "neighbour", the authors have kept the sentences describing deletions. I still think that the definition is not correct: They do not want that the neighbours of x become the neighbours of y, but want to add them to the existing neighbours of y. Simply deleting the word "the" would be enough.
- The typography used in the algorithms could be improved. Also notice that "bipartitions1" is later refered to as "bipartitions_1". This comment about typography also applies to the operations "Del" and "Ins" in Definition 1.
Reviewed by Barbara Holland, 18 Nov 2020
The authors have addressed all the comments I had on the earlier manuscript.
I will leave the other reviewers to assess if the proofs are now satisfactory as they were the ones who spotted some extra issues.
https://doi.org/10.24072/pci.mcb.100002.rev23Evaluation round #1
DOI or URL of the preprint: 10.1101/2020.09.14.293522
Author's Reply, 11 Nov 2020
Dear Editor,
Thank you for your and the reviewers' detailed feedback on our submission.
We have addressed all the points in this revised version of the manuscript.
Please find attached a letter with our point-by-point replies to the reviewers' comments, and our revised manuscript with changes highlighted.
Best regards,
Nadia El-Mabrouk
Decision by Céline Scornavacca , posted 07 Oct 2020
, posted 07 Oct 2020
Dear authors,
The paper has been reviewed by four (!) reviewers, who did a tremendous job, reading the paper in depth and providing constructive comments.
They all agree that the paper is generally well-written and that deserves publication after fixing some issues that makes it more complex than needed (e.g. the rooted vs unrooted issue, some missing details in the proofs, a better explanation of the algorithm and of the purpose of Section 5.1, ...).
When preparing the revision, the authors should answer to the major points of each reviewer in a separate text and provide a file where the modifications are highlighted (e.g. using difflatex). Also, they should compile the paper using an "plain" latex template (no Bioinformatics logo, please) and put the paper on a preprint server (e.g. arxiv).
Thanks again for your submission, and I look forward to reading the revised version.
Best regards,
Céline Scornavacca
Reviewed by Barbara Holland, 30 Sep 2020
Reviewed by Gabriel Cardona, 07 Sep 2020
The manuscript under review defines what the authors call the Labeled Robinson-Foulds Distance, and give a linear algorithm for its computation.
The paper is well written and (mostly, see below) technically sound. The results are of interests in the area of phylogenetics.
In my opinion, there are a few "major" issues that I think the authors should address, and I also have some other "minor" comments and suggestions.
Major:
- Generic: In my opinion, labeled trees usually refer to trees whose nodes are uniquely labeled by a given set. Moreover, in the context of generalizing the RF distance, bipartitions admit an straightforward generalization to (uniquely) labeled trees, even when not all internal nodes are labeled. Hence, when reading the title of the paper, my first thought (and I guess that also that of many other potential readers) was this particular generalization. Therefore, I'd suggest using a different name.
- P1,top: The authors should remove the Bioinformatics/Oxford logo in order to be published in PCI.
- P3,C1,par 3: "can then be deduced from the RF distance of the "unrooted version"": This depens on how you define the "unrooted version" of a tree. With your notations it should mean to forget the root and eliminate it if it has degree two. Two different rooted trees (hence at distance >0) may have the same unrooted version (hence at distance 0). Therefore, the "rooted version" of the distance cannot be deduced from the "unrooted version".
- P2,C1,par 4: "In this paper, we focus on unrooted trees, thus avoiding the special case of the root. Therefore, from now on, all trees are considered unrooted.": One of the things that sometimes makes this manuscript a little hard to read is when they try to write all definitions and results in Section 2 suitable for both rooted and unrooted trees. If in the end they only consider unrooted trees, I'd suggest to make this Section 2 more specific to unrooted trees. Note, however, that islands, defined below, are rooted if I understood it correctly (or they allow for nodes of degree two).
- P4,C1,definition 3: I find this definition more intricate than needed. I'd say that islands are exactly the connected components obtained by removing the internal good edges such that they contain at least a leaf of the original tree. Also, with the given definition it is not clear if a node all whose incident edges are good internal edges constitutes by its own an island (but the definition allows for it).
- P4,C2,top: Please give a precise reference for the Lemma. Also, I see this formulation too intricate: why not simply say that the partition on the set of leaves induced by the islands is the same in both trees?
- P5,C1,par 5: "we clearly require at least ε(I) + ε(I') node removals": I think it is true, but more details should be given: it must use that all edges are bad, for instance.
- P5,C2,par 2: "This sequence of operations then leads to the tree T'j, which is the same as Tj except possibly the two labels of x and y": This should be proved.
- P5,C2,par 5: "and thus P can be reordered in the form...": It should be justified that operations can be reordered.
- P5,C1,par 2,3,4:
- The three paragraphs should be rewritten and expanded. Notice that the purpose of the manuscript is giving a linear-time algorithm. Hence, all these steps of the algorithm have to be fully explained, including the proofs of the running times of each step.
- Also, there is in my opinion a problem on how the iteration is done (in terms of good edges): First, some edges may not adjacent to islands, and if they are, it has not been mentioned that this islands are unique (as the pseudocode assumes). I'd suggest iterating over islands instead of good edges; it would also avoid the problem of having to check if an island has been visited or not.
- P5,C1,algorithm: There is a strange mixture of lines with extremely detailed pseudocode and other ones too vague. For instance, although it may be clear what from the context what getBipartitions, getIslands, islandPair do, it should at least be explicitly stated. Maybe it makes no sense to write it as a latex algorithm. The same information can be given with an itemize (nested, if needed), so that more details can be given on what is exactly computed, combining the information in the algorith with the description given in the rest of the section.
Minor:
- P2,C2,par 2:
- "admits a single,..." sounds strange: it is a tree with a distinguished node called its root
- "Now an internal node x is binary": specify that internal and different from the root.
- P2,C2,par 3: "y is a descendant of x if y is on the path...": It is easier to say that the path from r to y passes through x
- P2,C2,par 7: "As recalled in Briand et al...": Please give the original reference (where it was first proved).
- P2,C2, par -3:
- "become the children...": Seems as if the children of y were replaced by those of x; in fact, the children of x (except for y) are added to the children of y
- "Del(T, x, y)": The "Del" looks bad (typographically). It should be an \operatorname or \DeclareMathOperator. Same for the other operations.
- "removing the edge {x, z}...": Not needed; when you remove x, all these edges are removed by the definition of node removal.
- P3,C1,par 2: "In the case of rooted trees, the RF distance is defined as the symmetric difference between the clades of the two trees.": Repeated (appears above)
- P3,C1,par 3: "The only thing that can make bipartitions and clades differ in number is rooting into a bad edge.": I don't understand what this sentence means.
- P3,C2,proof of Lemma 1: It is an edit distance. Only the reversibility of the operations has to be remarked; the rest is a classical result.
- P3,C2,proof of Lemma 2: Maybe I miss some subtle detail, but I see this result as straightforward: since there is a single label, the operation Sub cannot be applied.
- P3,C2,par -4: "Edge contraction Cont(T, {x, y}) similar to...": Similar or equal?
- P3,C2,par -3: "Node flip Flip(x, λ)": The other operations have T as their first argument.
- P4,C1,lemma 3: I'd suggest giving an example where the inequality is strict.
- P4,C1,definition 3: "..., and all terminal edges of I are good edges of T.": I think this condition is not needed, since all terminal edges are good edges.
- P4,C1,par -1: " while each good edge belongs to exactly two islands of T". I'd say that it belongs to no island at all, but maybe this "belongs" is defined to make it happen. In any case, it should be clarified. See also my objection on the definition of island above and notice that the caption of Fig. 2 also uses this notion of "belongs".
- P5,C2,lemma 6: It should be stated in terms of node deletions, or else define what it means here the deletion of an edge.
- P5,C2,par 2:
- "...a path transforming T intoT ." -> "... a path transforming T into T and assume that it involves the deletion of a good edge."
- In the description of cases (1)...(3): Why not do it the other way?: start with the concrete cases (2),(3) and then say "otherwise o'k=ok"
- Clarify what "does not affect node y" and "rename z as x" means.
- P5,C2,par 5: "As islands can only share good edges, ...": It should be stated what it means.
- P6,C1,par 3:
- "stem" -> "steps"
- "Using the node-to-island map": This map has not been defined in the algorithm.
- P6,C1,par 4: "is implemented by adding" -> "is updated by adding"
- P6, section 5.1: I don't see the relevance of this experiment.
- P6,C2,par 1: "Finally, we showed that the new distance is computable for an arbitrary number of label types associated with internal nodes of the tree." I don't understand this sentence. I'd suggest modifying the previous sentence "...being a metric and reducing to..." -> "...being a metric, even for an arbitrary number of labels, and reducing to..."
- P6,C2,par -3: I don't see why "Our experimental results illustrate the utility of computing tree distances taking labels into account, as the conventional RF distance is blind to label changes." It is obvious that RF distance does not take labels into account, and it is independent of any experimental result.
Reviewed by Jean-Baka Domelevo Entfellner , 28 Sep 2020
Summary of the paper
The authors of the manuscript titled "A Linear Time Solution to the Labeled Robinson-Foulds Distance Problem" define an new formulation of labeled Robinson-Foulds (LRF) distance on labeled phylogenetic trees. This formulation, based on operations on nodes, is an alternative to some other formulation (based on edge edits) of the LRF that they introduced in another paper to be published this year. This new formulation enables exact computation of the distance in linear time with respect to the number of leaves, and the authors describe an algorithm they implemented in Python. The authors conclude the paper with computation on some test tree and comparisons between the RF, the approximate LRF (from their earlier paper) and the exact LRF. They also demonstrate, based on simulated trees, that denser taxon sampling seems to improve the accuracy of the inference of a labeled gene tree, accuracy which they measure in terms of LRF distance between true and inferred tree.
Scope and importance of the findings
Though it is affected by a number of shortcomings, the Robinson-Foulds (RF) distance is still widely used on unlabeled phylogenetic trees. This is due to the fact that it is a true metric, and that it is easy to interpret and to calculate, in sub-linear time. The LRF is the extension of the RF distance to labeled trees, and it derives directly from the Tree Edit Distance (Zhang et al, 1989). It is important to mention (and so the authors do) that a seizable amount of literature has already been published on the Tree Edit Distance (TED), which is actually a generalization of the LRF to either ordered or unordered labeled trees with a cost function on node edit operations. Yet, since the TED has been used so far mostly in other data science fields (hierarchical clustering on various types of data, grammar parsing, RNA secondary structure analysis, etc), the TED (end therefore the LRF) has not so far received much attention from the phylogenetics community. The LRF being the equivalent of the widely-used RF distance for the world of labeled trees (where node labels serve e.g. to discriminate between speciation and duplication nodes), one can guess that the LRF distance could be more widely used in the coming years, provided that it is well established and easily computable. To this end, the contribution of the submitted paper is unquestionable, and I am in favour of its publication. Nevertheless, I identified a number of shortcomings in the paper, that the authors should address before publication. I list them below. Most of these remarks and suggestions are also featuring in my handwritten comments on the manuscript whose scan I attach herein. I invite the authors to read all the comments and edits I suggest on the scanned copy of the paper.
Remarks and suggestions to be addressed by the authors.
Though the authors duly cite the body of literature relative to the TED distance, including the 1989 paper by Zhang and Shasha and the 1992 paper by Zhang, Statman and Shasha, they don't explain why a linear solution is demonstrable for the calculation of the LRF edit distance on phylogenetic trees, while the 1992 reference showed that the TED on unordered labeled trees (trees in which the neighbours of a node constitute an unordered set) is NP-complete. This is because the literature on TED considers a non-constant cost function on edit operations, while for the RF and LRF distances, every operation has constant unitary cost. This should be made explicit in the paper.
Similarly, as not all the readers will be familiar with the formulation of the RF distance as a total number of alpha (edge or node deletion) and alpha^(-1) (edge or node insertion) operations on a transformation path, it would be good to say a few precise words about it in section 2.1, for instance when the authors say "The Robinson-Foulds or Edit distance [...] is the length of a shortest path of node edit operations transforming [...]": the authors should state clearly which were the edit operaions originally devised by Robinson and Foulds in their 1981 paper.
There is a bit of a confusing or uncomfortable back-and-forth between rooted and unordered trees at the beginning of the paper, mainly for historical reasons (the TED edit distance having been used in communities, like the one of classification, where trees are naturally rooted trees). This lasts until the end of section 2, when the authors say "Therefore, from now on, all trees are considered unrooted." And yet, in the following sections, the authors keep using the words "child" and "children", while they should have used the word "neighbour(s)". Sticking to the vocables of rooted trees while talking of unrooted trees may confuse the reader.
Throughout the paper, the authors seem to have turned a blind eye to the fact that unrooted trees may still contain nodes of degree 2, while the whole algorithmic machinery developed in the paper CANNOT accommodate such trees. Accepting trees in which at least one internal node has degree 2 gives birth to situations in which, according to the definition of the node edit operations by the authors in the present paper, there exists a transformation path from T to T' but NOT from T' to T, which obviously annihilates the proof that makes LRF a metric. Although internal nodes of degree 2 are usually not seen as valid phylogenetic trees, for the sake of mathematical accuracy, the authors should mention somewhere that they consider only trees whose intenal nodes all have degree 3 or greater.
The authors should be careful, when they define subtrees in the fourth paragraph of section 2, to pay attention that their current definition, as written in the version I reviewed, does not preserve connexity.
In several occurrences, the authors talk about the symmetric difference between sets A and B while they actually mean the size of that difference (i.e. the number of elements in the union of A-B and B-A).
In definition 3 (section 3.1), please pay attention to the fact that, contrary to what the authors wrote, islands do not form a partition of the tree: any two islands share a good edge and its two connected nodes.
Lemmas 1, 2, 3, 5 and 6 all come with their proofs in this paper. In contrast, the proof for lemma 4 does not feature here. This is puzzling, and the reader needs to go to Briand and al (2020) to find the proof (?). In case it wasn't included here for the sake of brevety, please mention this fact, together with a few words describing a rough summary of the proof.
In the second paragraph of the proof for lemma 5, when the authors say: "On the other hand, since an edit operation can remove or insert at most one edge, [...] we clearly require [...]", they should rather say that the grounding for that part of the proof comes from the fact that all internal nodes in an island are bad ones, and therefore need to be removed.
Please write "label-disjoint" everywhere, altering the few occurrences where "label disjoint" is written without hyphenation.
The proof of lemma 6 is quite difficult to follow and it leaves the reader under the impression that weaknesses exist in there that are not addressed by the authors. The proof relies on the construction of an alternative sequence of edit operations transforming T into T', and it makes assumptions whose validity it is uneasy to check. For instance, where is the guarantee that at that stage, in the tree T_{k-1}, those z and x nodes will be neighbours? That is not straightforward, and in my opinion, the proof needs a bit of reworking or rewriting to clarify this point.
In the proof for lemma 6, B1 and B2 are defined as subsets of leaves (taxa), but are used as subtrees. Although every reader will understand your point there, please clarify this for the sake of mathematical correctedness.
In the logical description of the algorithm (section 4), on line 8 of the pseudocode, (x1, y1) and (x2, y2) are ill-defined; we understand each of those four elements denotes an island, but we don't understand why pairs of islands whould be examined in pairs. The text should explain this (more) clearly.
In Figure 5, it would be informative to display in the top graph (relative to the RF distance) the line with equation y = 0.7*x, since an average 30% of the random edit operations are node label substitutions, to which the RF distance is totally blind. In that sense, the line with equation y = x is not the "expected"/"fair" regression line here.
Reviewed by anonymous reviewer 1, 28 Sep 2020
Summary of the paper and its contribution
The authors introduce a new distance (LRF-distance) for labelled trees, where not only leaves (as in 'classical' phylogenetic trees), but also internal nodes are labelled. This is of particular interest in gene trees, where internal nodes are labelled by specific evolutionary events they represent, e.g. speciation and duplication. The metric introduced in this paper is based on the Robinson-Foulds distance, which is not defined for trees where internal nodes are labels, and for unlabelled trees the LRF-distance actually equals the Robinson-Foulds distance. An algorithm for computing the LRF-distance in linear time is presented as well as a simulation study to compare the LRF-distance to other distance measures on labelled trees and another simulation study to evaluate the benefits of dense taxon sampling.
General Impression
The paper is well written and does not only introduce a new metric for labelled trees, but also presents an algorithm to compute this distance in linear time. It moreover contains a simulation to study the effect of denser taxon sampling on tree inference of labelled gene trees, which uses this new metric. There however are a few things that can and should be improved, but most of them are technical and do not influence the quality of the results of this paper. Among a few minor things (see the list below), I discovered the following issues:
It is not clear why RF is introduced for rooted trees in Section 2.1. and it is not always clear where the authors refer to rooted and where to unrooted trees. As the main focus of the paper is on unrooteed trees, I recommend only talking about such trees and not about rooted trees, unless the results of the paper can equally be applied to rooted trees, in which case this should be mentioned.
In the proof of Thm 1 it is not necessary to show that the order of Pi can be changed on the path P. It is sufficient to show that all Pi are shortest paths and there is a shortest path preserving all good internal edges.
To me it is not clear what the aim of Section 5.1 is. Robinson Foulds, ELRF, and LRF are compared by taking trees, randomly performing edit operations that define LRF (or ELRF) and then computing RF, ELRF, and LRF distance between the computed trees. The results are as one would expect -- which is resulting from the fact that for ELRF and LRF the corresponding edit operations have been used while for RF not only the operations corresponding to RF have been used. It hence is not obvious to me what the purpose of this empirical comparison of RF, ELRF, and LRF is.
Further Comments
- Def 1, Node deletion: not only the edge {x,z}, but also the edge {x,y} should be deleted in Del(T,x,y)
- Def 2, Node label substitution: It is not clear of the label lambda of x can be replaced by the same label lambda by such a move
- Paragraph after Def 2: 'The two following lemma state [...]' -> should be 'lemmas'
- Proof Lemma 2, Paragraph 2: 'Conversely, Let P be a path labeled node edit [...]' -> should be 'Conversely, let P be a path of labeled node edit [...]'
- Paragraph after Def 3: 'start tree' -> should be 'star tree'
- same paragraph: good internal edges belong to exactly two islands - from my understanding only the endpoints belong to islands, the edges itself are not in any of the islands
- Fig 2: It is very hard to see what the islands of these trees are and the caption does not help there
- Fig 4: There are a number of leaf labels missing and the number of inversions between the two topmost trees on the right is wrong (should be 2 instead of 3)
- Proof Lemma 6, 3rd line: first Bx and By are used, then B1 and B2
- Proof Thm 1: 3rd line: 'islands share good edges' -- it is not clear what that means










